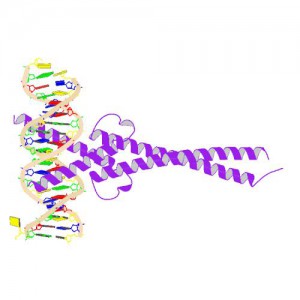
In 1982, picked up because of its homology to chicken virus genes that could transform cells, MYC became one of the first human genes identified that could drive cellular transformation (1,2). Since that time countless laboratories have prodded and poked the human MYC gene, the MYC protein, their homologs in other animal models, and their transforming viral counterparts.
MYC is a transcription factor and forms heterodimers with a required protein partner, MAX, before binding to the E box sequences of DNA regulatory regions (3). MYC regulates gene expression of many targets through interactions with a host of proteins, often referred to as the MYC Interactome (2). In fact, MYC is estimated to bind 10–15% of the genome, and it regulates the expression of genes that are transcribed by by each of the three RNA polymerases (2).
MYC plays a central role in regulating cell growth, proliferation, apoptosis, differentiation and transformation, acting as a central integrator of cellular signals. MYC is tightly regulated at multiple levels from gene expression to protein stability. Dysregulation (usually upregulation) of the amount and stability of Myc protein is observed in many human cancers. Even in cancers in which MYC is not directly involved in transforming cells, its normal expression is often required to support the extracellular matrix and/or vascularization necessary for tumor growth and formation (4).
Because MYC is such a central player cancer pathology, it is an attractive target for cancer therapeutics (2) .
But while MYC is central in pathology, it also is central in biology, and scientists have noted that, with the exception of Burkitt’s lymphoma (2), MYC is an under mutated gene. This under mutation and the typical upregulation and amplification in many cancers suggests that MYC function may be required for proper biological function and development, raising concerns about off-target effects if MYC function is reduced or eliminated as a result of therapeutic activity. So far, though, genetic silencing of MYC has only resulted in mild side effects that were reversed when expression was restored (3).
A second factor that complicates therapeutics targeted against MYC is that it encodes an intrinsically disordered protein with no enzymatic activity; therefore there is no active site or catalytic domain to easily block or inhibit with a small molecule. Furthermore, MYC is only functional after interacting with its protein partner MAX to form a heterodimer in which MYC contributes the transactivation domain.
MYC interacts with many proteins including transcriptional activators, transcription elongation factors, proteins that are involved in transcriptional repression, and proteins involved in regulating protein stability (3).
Disrupting these protein:protein interactions of MYC and its many partners is one approach to targeting MYC in cancer. One group has demonstrated proof-of-concept in targeting MYC:MAX interactions by inducible expression in mice of a dominant interfering helix-loop-helix zip dimerization domain, called Omomyc in mice. Omomyc was effective in treating KRas-driven lung cancer and had minimal off-target effects (5,6). Two other groups have demonstrated that a small molecule inhibitor, JQ1, to inhibit the acetyl-lysine recognition domains (bromodomains) of MYC partner proteins that are involved in transcription initiation and elongation can downregulate MYC activity (7,8). These studies would suggest then that targeting MYC protein:protein interactions could be a successful strategy for designing successful, safe cancer therapeutics.
However MYC, as noted above, interacts with 10-15% of the genome, and very little is known about the nature and biology of most of these interactions. Proteome-wide, big data screens can yield information about the abundance of proteins, but neither they nor more traditional methods of studying protein:protein interactions can provide understanding about what is happening inside the cell in a physiological system. To understand the biology of each protein:protein interaction and what happens when it is disrupted in a cellular system, techniques that can interrogate the individual interactions inside cells are needed.
One such technique is BRET (bioluminescence resonance energy transfer). With BRET, energy is transferred between a bioluminescent donor to a fluorescent receptor. Because this energy transfer is resonance energy transfer through dipole coupling, the two molecules to which the donor and receptor molecules are attached must be less than 10nm apart and in the correct orientation (9). These requirements mean that the signal received from BRET analysis is highly specific to the pair of proteins or (protein and ligand) being studied.
When performing BRET analyses, the donor and receptor molecules that are attached as reporters to the proteins of interests need to be small so that they are least likely to cause steric hindrances that will disrupt natural interactions. Using a small luciferase molecule like NanoLuc® Luciferase helps to address this concern, and because the NanoLuc® Luciferase is so bright, it can be expressed at low (more like endogenous) levels with enough energy remaining for the resonance energy transfer to the fluorescent receptor.
Perhaps with the new tools to interrogate the specific protein:protein interactions of MYC and its many, many partners, we will be able to more fully understand the biology of that is disrupted in cancers that are reliant upon MYC overexpression and approach our research and development of therapeutics targeted toward MYC with forethought and creativity.
Literature Cited
- Varmus, H.E. (1984) The molecular genetics of cellular oncogenes. Rev. Genet. 18, 553–612.
- Tu, W.B. et al. (2015) Myc and its interactors take shape. Biophys. Acta 1849, 469–83.
- Jung, K-Y. et al. (2015) Perturbation of the c-Myc-Max Protein-Protein Interaction via synthetic alpha helix mimetics. Med. Chem. 58, 3002–24.
- Fletcher, S. and Prochownick, E.V. (2015) Small-molecule inhibitors of the Myc oncoprotein. Biophys. Acta 1849, 525–43.
- Soucek, L. et al. (2008) Modelling Myc inhibition as a cancer therapy. Nature 455, 679–83.
- Soucek, L. et al. (2013) Inhibition of Myc family proteins eradicates KRase-driven lung cancer in mice. Genes and Dev. 27, 504–13.
- Delmore, J.E. et al. (2011) BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146, 904–17.
- Zuber, J. et al. (2011) RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukemia. Nature. 478, 524–8.
- Rickey, T. (2015) Studying protein:protein interactions with BRET and NanoLuc° Luciferase. Promega PubHub [Internet http://www.promega.com/resources/pubhub/features/bret-nanoluc-luciferase-and-protein-protein-interactions/ Accessed May 7, 2015 ]
Michele Arduengo
Latest posts by Michele Arduengo (see all)
- An Unexpected Role for RNA Methylation in Mitosis Leads to New Understanding of Neurodevelopmental Disorders - March 27, 2025
- Unlocking the Secrets of ADP-Ribosylation with Arg-C Ultra Protease, a Key Enzyme for Studying Ester-Linked Protein Modifications - November 13, 2024
- Exploring the Respiratory Virus Landscape: Pre-Pandemic Data and Pandemic Preparedness - October 29, 2024