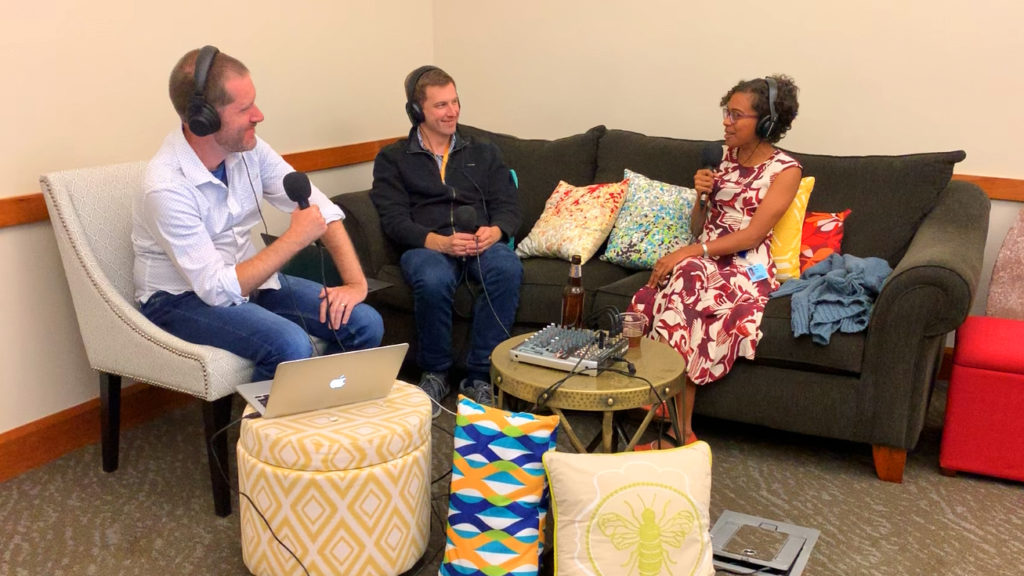
This past weekend was the 9th Annual Wisconsin Science Festival, and we at Promega were excited to join in the celebration of science throughout the state. We participated in the Discovery Expo on Thursday and Friday, where dozens of demonstrations and exhibits were scattered throughout the Wisconsin Institute for Discovery building. Thousands of children on field trips filled the halls, eager to poke and prod at strange and exciting new things.
At our table, we talked about the science of bioluminescence. With 3D-printed firefly luciferase models in hand, we showed the glow of recombinant luciferase to the incoming children and explained to them how scientists could use bioluminescence like a tiny “flashlight” to look inside of cells and watch what’s happening. Our learners received a nice little reward for their attentiveness in the form of glow-in-the-dark firefly stickers.

 I used to love taking magazine quizzes to learn more about myself. I thought it would be fun to create a quiz to help you find out what scientific career path may be the best fit for you. Be open-minded while taking the quiz and remember that this is just for fun!
I used to love taking magazine quizzes to learn more about myself. I thought it would be fun to create a quiz to help you find out what scientific career path may be the best fit for you. Be open-minded while taking the quiz and remember that this is just for fun!