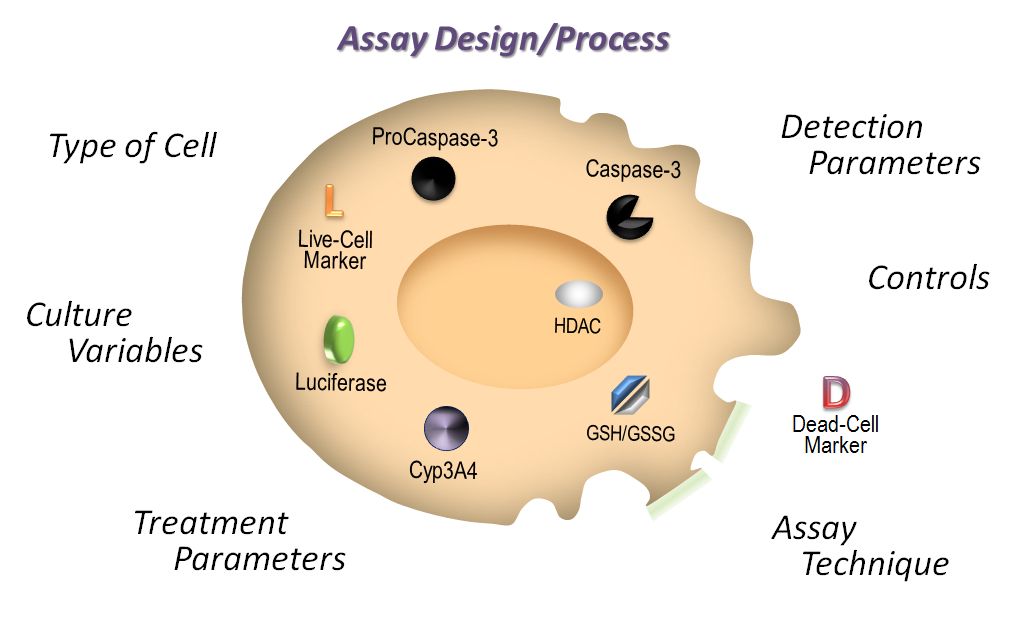
This is the second in a series of blog posts covering topics to consider when designing and performing cell-based assays. In the first installment, we discussed the importance of choosing the right cell type for your assay. Here we will discuss how cell culture conditions affect cell-based assays.
Continue reading “Considerations for Successful Cell Based Assays II: Cell Culture Conditions”Tips and Tools
Fixed in the Past, Focus on the Future
“I would do more with my samples, but it’s just not possible…I know there’s probably a wealth of information in there, but there is just no way to get it out…I’ve got blocks of tissue sitting in the lab, experiments I want to run, but no good way to get clean nucleic acids out.”
These are a few of the comments I heard when talking with scientists at the American Society of Human Genetics meeting last week in Montreal. They, and countless other researchers, are sitting on a treasure trove of information that may have been locked away a few months ago, a few years ago, or decades ago. I’m referring to formalin-fixed, paraffin-embedded (FFPE) tissue blocks. It is estimated that there are upwards of 400 million tissue blocks archived globally and scientists are clamoring to find ways to best utilize nucleic acids derived from these tissues in applications like qPCR, microarrays, and next generation sequencing.1 Continue reading “Fixed in the Past, Focus on the Future”
Optimized Protein Expression: Flexi Rabbit Reticulocyte Lysate
mRNAs commonly exhibit differing salt requirements for optimal translation. Small variations in salt concentration can lead to dramatic differences in translation efficiency. The Flexi® Rabbit Reticulocyte Lysate System allows translation reactions to be optimized for a wide range of parameters, including
Mg2+ and K+ concentrations and the choice of adding DTT. To help optimize Mg2+ for a specific message, the endogenous Mg2+ concentration of each lysate batch is stated in the product information included with this product.
The following references utilize the features of Flexi Rabbit Reticulocyte Lysate System to investigate certain parameters of translation:
Vallejos, M. et al. (2010)The 5′-untranslated region of the mouse mammary tumor virus mRNA exhibits cap-independent translation initiation. Nucl Acids Res. 38, 618–32. Identification of internal ribosomal ribosomal entry site in the 5’ untranslated region of the mouse mammary tumor virus mRNA.
Spriggs, K. et al. (2009) The human insulin receptor mRNA contains a functional internal ribosome entry segment. Nucl. Acids. Res. 17, 5881–93. Identification of a functional internal ribosome entry site in the human insulin receptor mRNA.
Powell, M. et al. (2008) Characterization of the termination-reinitiation strategy employed in the expression of influenza B virus BM2 protein. RNA 14, 2394–06. Analysis of the mRNA signals involved in the expression of influenza B virus BM2 protein.
Sato, V. et al. (2007) Measles virus N protein inhibits host translation by binding to eIF3-p40. J. Vir. 81, 11569–76. Charaterized the effect of the measles virus N protein binding to the translation initiation factor eIF3-p40 on the expression of various proteins in rabbit reticulocyte lysate.
Hirao, K. et al. (2006) EDEM3, a soluble EDEM homolog, enhances glycoprotein endoplasmic reticulum-associated degradation and mannose trimming. J. Biol. Chem. 281, 9650–58. The EDEM3 protein was expressed in the presence of canine microsomal membranes to establish that co-translational translocation occurs into the endoplasmic reticulum.
Shenvi, C. et al. (2005) Accessibility of 18S rRNA in human 40S subunits and 80S ribosomes at physiological magnesium ion concentrations–implications for the study of ribosome dynamics. RNA 11, 1898–08. Characterization of ribosome dynamics under different ionic conditions.
Nair, A. et al. (2005) Regulation of luteinizing hormone receptor expression: evidence of translational suppression in vitro by a hormonally regulated mRNA-binding protein and its endogenous association with luteinizing hormone receptor mRNA in the ovary. J. Biol. Chem. 280, 42809–16. Examined the affect of luteinizing hormone receptor mRNA binding protein on transltional suppression of luteinizing hormone receptor RNA.
Optimization of Western Blots Detecting Proteins Synthesized Using Cell-Free Expression #2
Detection of protein expressed using cell-free systems is required for most applications such as protein:protein interaction and protein:nucleic acid interaction studies. Traditionally, one adds radioactive [35S]methionine to cell-free expression reactions, and the methionine is incorporated into the expressed protein, allowing detection by autoradiography. Many researchers are moving away from radioactivity. Traditional Western blot analysis provides the researcher a nonradioactive method for detection but, if performed improperly, can result in high background, which can mask expressed proteins and affect downstream applications.
One critical step in producing low-background, high-signal Western blots is choosing the correct dilution of the primary antibody. Typically the manufacturer recommends antibody dilution from 1:1,000 to 1:2,500 for standard western blotting experiments. However when using crude lysates as a source of the target protein, these recommendations exhibit significant background. When the antibody was diluted 1:50,000, background was decreased significantly, and the positive signal was a large percentage of the total signal.
As a general recommendation when performing Western blot analysis of proteins expressed in cell-free systems, one must experimentally determine the optimal dilution of the primary antibody. In the Western blots performed in this study, primary antibodies were diluted ~50-fold more than the provider’s recommended dilution.
For additional technical details refer to this recent article published in Promega’s PubHub:
Hook, B and Schagat, T. (2011) Non-Radioactive Detection of Proteins Expressed in Cell-Free Expression Systems Promega Corporation Web site. Accessed August 17, 2011.
Troubleshooting T-Vector Cloning
Why do few of my pGEM®-T or pGEM®-T Easy Vector clones contain the PCR product of interest?
There are several possible reasons why the PCR product may not be recovered after ligation, bacterial transformation and plating when using the pGEM®-T or pGEM®-T Easy Vector Systems.
The PCR fragment may not be A-tailed. Without the A overhangs, the PCR product cannot be ligated into a T vector. Use a nonproofreading DNA polymerase like GoTaq® DNA Polymerase for PCR. If a proofreading DNA polymerase is used, A overhangs will need to be added. Purify the PCR fragment, and set up an A-tailing reaction (see the pGEM®-T and pGEM®-T Easy Vector Systems Technical Manual #TM042). The A-tailed product can be added directly to the ligation as described in the pGEM®-T or pGEM®-T Easy Vector protocol.
The insert:vector ratio may not be optimal. The ideal ratio for each insert to a vector can vary. For example, the Control Insert DNA works well at a 1:1 ratio, but another insert may be ligated more efficiently at a 3:1 ratio. Check the integrity and quantity of your PCR fragment by gel analysis. Optimize the insert:vector ratio (see Technical Manual #TM042).
Multiple PCR products were amplified and cloned into the pGEM®-T or pGEM®-T Easy Vector. Other amplification products including primer dimers will compete for ligation into the T vector, decreasing the possibility that the desired insert will be cloned. To minimize other competing products, gel purify the PCR fragment of interest.
Promega Technical Services Scientists are here to assist you in troubleshooting your experiments at any time. Contact Technical Services.
Related Posts
Tips for Multiplex Cell-Based Assay Success
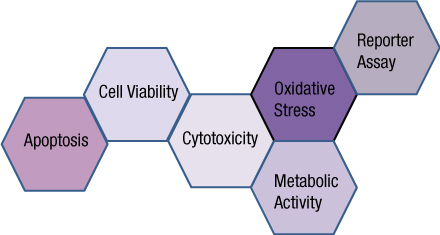
Analyzing more than one cellular biomarker (multiplexing) in a single sample is advantageous for a number of reasons. Multiplexing allows researchers to save money and time, while conserving critical samples. In addition, understanding the relationship between cell biomarkers can provide a more complete picture of cell health, leading to improved predictive models for drug discovery. Understanding biomarker relationships can also minimize ambiguity in the data set and validate if a treatment effect is real or an artifact of the system. To avoid repeat experiments and extract the most physiologically relevant data from multiplex cell-based assays, we discuss considerations around assay choices, cell type, cell culture, treatment parameters, detection and appropriate experimental controls.
Continue reading “Tips for Multiplex Cell-Based Assay Success”Screening for Protein Activity Using Cell-Free Expression
The analysis of functional protein typically requires lengthy laborious cell based protein expression that can be complicated by the lack of stability or solubility of the purified protein. Cell free protein expression eliminates the requirement for cell culture thus providing quick access to the protein of interest (1).
The HaloTag® Technology provides efficient, covalent and oriented protein immobilization of the fusion protein to solid surfaces (2).
A recent publication demonstrated the feasibility of using cell free expression and the HaloTag technology to express and capture a fusion protein for the rapid screening of protein kinase activity (3). The catalytic subunit of human cAMP dependent protein kinase was expressed in a variety of cell free expression formats as a HaloTag fusion protein. The immobilized cPKA fusion protein was assayed directly on magnetic beads in the active form and was shown to be inhibited by known PKA inhibitory compounds.
Therefore this unique combination of protein expression and capture technologies can greatly facilitate the process of activity screening and characterization of potential inhibitors
- Zhao, K.Q. et al. (2007) Functional protein expression from a DNA based wheat germ cell-free system. J. Struc. Funct. Genomics. 8, 199-208.
- Los, G.V. and Wood, K. (2007) The HaloTag: A novel technology for cell imaging and protein analysis. Meth. Mol. Biol. 356, 195-208
- Leippe DM, Zhao KQ, Hsiao K, & Slater MR (2010). Cell-free expression of protein kinase a for rapid activity assays. Analytical chemistry insights, 5, 25-36 PMID: 20520741
Protease K Protection Assay: Cell Free Expression Application
Microsomal vesicles are used to study cotranslational and initial posttranslational processing of proteins. Processing events such as signal peptide cleavage, membrane insertion, translocation and core glycosylation can be examined by the transcription/translation of the appropriate DNA in the TNT® Lysate Systems when used with microsomal membranes.
The most general assay for translocation makes use of the protection afforded the translocated domain by the lipid bilayer of the microsomal membrane. In this assay protein domains are judged to be translocated if they are observed to be protected from exogenously added protease. To confirm that protection is due to the lipid bilayer addition of 0.1% non-ionic detergent (such as Triton® X-100) solubilizes the membrane and restores susceptibility to the protease.
Are you looking for proteases to use in your research?
Explore our portfolio of proteases today.
Many proteases have proven useful for monitoring translocation in this fashion including Protease K or Trypsin.
The following are examples illustrating this application:
- Minn, I. et al. (2009) SUN-1 and ZYG-12, mediators of centrosome-nucleus attachment, are a functional SUN/KASH pair in Caenorhabditis elegans. Mol. Biol. Cell. 20, 4586–95.
- Padhan, K. et al. (2007) Severe acute respiratory syndrome coronavirus Orf3a protein interacts with caveolin. J.Gen.Virol. 88, 3067–77.
- Tews, B.A. et al. (2007) The pestivirus glycoprotein Erns is anchored in plane in the membrane via an amphipathic helix. J.Biol.Chem. 282, 32730–41.
- Pidasheva, S. et al. (2005) Impaired cotranslational processing of the calcium-sensing receptor due to signal peptide missense mutations in familial hypocalciuric hypercalcemia. Hum. Mol. Gen. 14, 1679–90.
- Smith, D. et al. (2002) Exogenous peptides delivered by ricin require processing by signal peptidase for transporter associated with antigen processing-independent MHC class I-restricted presentation. J. Immun. 169, 99–107.
6X His Protein Pulldowns: An Alternative to GST
![]() Pull-down assays probe interactions between a protein of interest that is expressed as fusion protein (e.g.,
Pull-down assays probe interactions between a protein of interest that is expressed as fusion protein (e.g.,
(e.g., bait) and the potential interacting partners (prey).
In a pull-down assay one protein partner is expressed as a fusion protein (e.g., bait protein) in E. coli and then immobilized using an affinity ligand specific for the fusion tag. The immobilized
bait protein can then be incubated with the prey protein. The source of the prey protein depends on whether the experiment is designed to confirm an interaction or to identify new interactions. After a series of wash steps, the entire complex can be eluted from the affinity support using SDS-PAGE loading buffer or by competitive analyte elution, then evaluated by SDS-PAGE.
Successful interactions can be detected by Western blotting with specific antibodies to both the prey and bait proteins, or measurement of radioactivity from a [35S] prey protein. bait) and potential interacting partners (prey).
The most commonly used method to generate a bait protein is expression as a fusion protein contain a GST (glutathione-S transferase) tag in E. coli. This is followed by immobilization on particles that contain reduced glutathione, which binds to the GST tag of the fusion protein. The primary advantage of a GST tag is that it can increase the solubility of insoluble or semi-soluble proteins expressed in E. coli.
Among fusion tags, His-tag is the most widely used and has several advantages including: 1) It’s small in size, which renders it less immunogenically active, and often it does not need to be removed from the purified protein for downstream applications; 2) There are a large number of commercial vectors available for expressing His-tagged proteins; 3) The tag may be placed at either the N or C terminus; 4) The interaction of the His-tag does not depend on the tag structure, making it possible to purify otherwise insoluble proteins using denaturing conditions. Continue reading “6X His Protein Pulldowns: An Alternative to GST”
Use of Multiple Proteases for Improved Protein Digestion
One of the approaches to identify proteins by mass spectrometry includes the separation of proteins by gel electrophoresis or liquid chromatography. Subsequently the proteins are cleaved with sequence-specific endoproteases. Following digestion the generated peptides are investigated by determination of molecular masses or specific sequence. For protein identification the experimentally obtained masses/sequences are compared with theoretical masses/sequences compiled in various databases.
Trypsin is the favored enzyme for this application, for the following reasons: A) the peptides contain a basic residue (Arg or Lys) on the C terminus and thus are good candidates for collision induced activation (CAD) in tandem experiments (low charge states and high mass-to-charge ratios); B) it is relatively Inexpensive; and C) optimal digestion conditions have been well characterized.
An inherent limitation of trypsin is the size of the peptides that it generates. For most organisms > 50% of tryptic peptides are less than 6 amino acids, too small for mass spectrometry based sequencing.
Are you looking for proteases to use in your research?
Explore our portfolio of proteases today.
One recent publication examined the use of multiple proteases (trypsin, LysC, ArgC , AspN and GluC) in combination with either CAD or electron-based fragmentation (ETD) to improve protein identification (1). Their results indicated a significant improvement from a single protease digestion (trypsin), which yielded 27,822 unique peptides corresponding to 3313 proteins. In contrast using a combination of proteases with either CAD or ETD fragmentation methods yielded 92,095 unique peptides mapping to 3908 proteins.
Swaney DL, Wenger CD, & Coon JJ (2010). Value of using multiple proteases for large-scale mass spectrometry-based proteomics. Journal of proteome research, 9 (3), 1323-9 PMID: 20113005