This the last in a series of four blogs about Quantitation for NGS is written by guest blogger Adam Blatter, Product Specialist in Integrated Solutions at Promega.
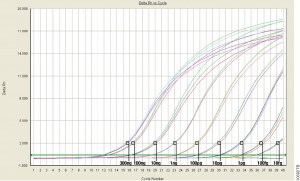
When it comes to nucleic acid quantitation, real-time or quantitative (qPCR) is considered the gold standard because of its unmatched performance in senstivity, specificity and accuracy. qPCR relies on thermal cycling, consisting of repeated cycles of heating an cooling for DNA melting and enzyamtic replication. Detection instrumentation is capable of measuring the accumulation of DNA product after each round of amplification in real time.
Because PCR amplifies specific regions of DNA, the method is highly sensitive, specific to DNA, and it can determine whether a sample is truly able to be amplified. Degraded DNA or free nucleotides, which might otherwise skew your quantiation, will not contribute to the signal, and your measurement will be more accurate.
However, while qPCR does provide technical advantages, the method requires special instrumentation, specialized reagents and is a more time-consuming process. In addition, you will probably need to optimize your qPCR assay for each of your targets to achieve your desired results.
Because of the added complexity and cost, qPCR is a technique suited for post-library quantitation when you need to know the exact amount of amplifiable, adapter-ligated DNA. PCR is the only method capable of specifically targeting these library constructs over other DNA that may be present. This specificity is important because accurate normalization is especially critical for producing even coverage in multiplex experiments where equimolar amounts of several libraries are added to a pooled sample. This normalization process is essential if your are screening for rare variants that might be lost in background and go undetected if underrepresented in a mixed pool.
Read Part 1: When Every Step Counts: Quantitation for NGS
Read Part 2: Nucleic Acid Quantitation by UV Absorbance: Not for NGS
Read Part 3: Fluorescence Dye-Based Quantitation: Sensitive and Specific for NGS Applications